Ecological Drivers of Bacterial Community Assembly in Synthetic Phycospheres
Fu, H., M. Uchimiya, J. Gore, and M. A. Moran
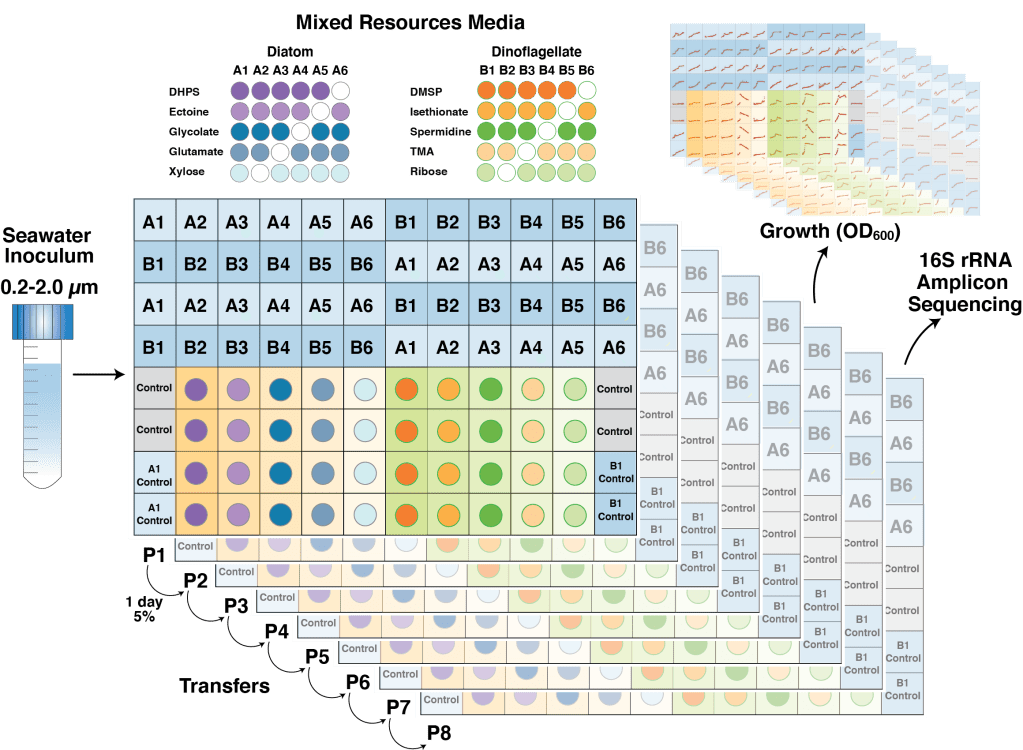
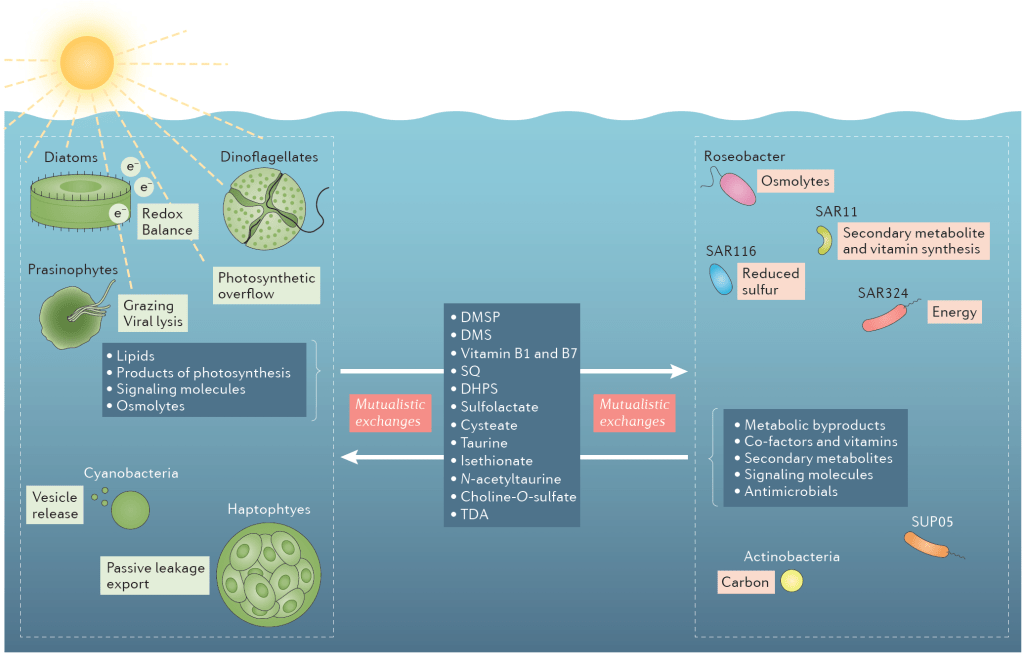
In the nutrient rich region surrounding marine phytoplankton cells, heterotrophic bacterioplankton transform a major fraction of recently fixed carbon through the uptake and catabolism of phytoplankton metabolites. We sought to understand the rules by which marine bacterial communities assemble in these nutrient-enhanced phycospheres, specifically addressing the role of host resources in driving community coalescence. Synthetic systems with varying combinations of known exometabolites of marine phytoplankton were inoculated with seawater bacterial assemblages, and communities were transferred daily to mimic the average duration of natural phycospheres. We found that bacterial community assembly was predictable from linear combinations of the taxa maintained on each individual metabolite in the mixture, weighted for the growth each supported. Deviations from this simple additive resource model were observed, but also attributed to resource-based factors via enhanced bacterial growth when host metabolites were available concurrently. The ability of photosynthetic hosts to shape bacterial associates through excreted metabolites represents a mechanism by which microbiomes with beneficial effects on host growth could be recruited. In the surface ocean, resource-based assembly of host-associated communities may underpin the evolution and maintenance of microbial interactions and determine the fate of a substantial portion of Earth’s primary production.